This week in PLOS Biology
In PLOS Biology this week, you can read about neural activity in bird song, the utility of mathematical models in evolutionary biology, how diet can shape the genome and the robustness of protein interaction networks.
Oiseaux Exotiques: How Birds Encode Birdsong
One of the fundamental problems in neuroscience is understanding the relationship between neural activity and the behaviour it produces. For motor systems, firing rates of neurons are traditionally quantified, whereas in sensory systems the temporal patterns in neuron spikes have emerged as critical. New work by Claire Tang, Samuel Sober, and colleagues in this issue of PLOS Biology indicates that, just as in sensory systems, the motor output that controls bird song is dictated not only by firing rate but also by the precise firing pattern and the information inherent in those patterns greatly exceeds the information available in simple spike counts. Read more in the accompanying synopsis.
Mathematical Models in Evolutionary Biology—What Are They For?
In a new Essay, Maria Servedio, Justin Yeh and co-authors discuss the utility of proof-of-concept modeling in evolutionary biology. They argue that the complexity of the processes and the long timescales involved in evolutionary biology means that mathematical approaches have long been required. However some people are still sceptical about the value of mathematical models in the field. They attempt to clarify, using examples, the unique utility of proof-of-concept models.
Codons May Depend on Diet

Ribosomes translate mRNA into protein using tRNAs, but the genetic code contains multiple synonyms – with the same amino acid specified by differing codon triplets. However all codons are not created equal – as some tRNAs read them with differing speed and accuracy. John Zaborske, Allan Drummond and colleagues have found that the way in which proteins are encoded has changed systematically across several closely related fruit fly species. Variation in the availability of a specific nutrient – queuine (from bacteria), influences tRNA anticodon modification from guanosine to queuosine in multiple drosophilid species (they test 12 species). The effects of this on translational accuracy have left their mark on the way that protein sequences are encoded in the flies’ genomes. Read more in the accompanying synopsis.
Signaling Networks Tolerate Domain Rearrangements
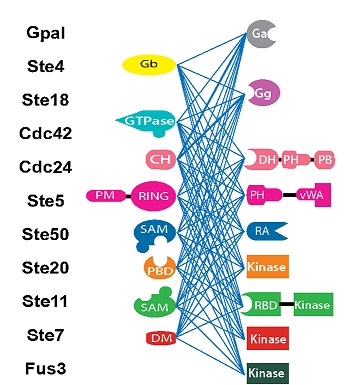
Cells use complex protein interaction networks to sense and process external signals. The proteins involved often comprise a string of several functional units called domains. Mutations that rearrange these domains therefore have the potential to create novels proteins that can process different signals – useful on an evolutionary timescale, but how well is this tolerated at the shorter timescale of the individual lifespan? Using a yeast model system, Paloma Sato, Sergio Peisajovich and colleagues found that signalling complexes are essentially more malleable than we previously thought. This is useful not only for evolution, but for manipulation of signaling pathways by synthetic biologists.

